File:Quarter_of_Arthrobacter_arilaitensis_Re117_genome.png
From Wikipedia, the free encyclopedia
Quarter_of_Arthrobacter_arilaitensis_Re117_genome.png (605 × 534 pixels, file size: 432 KB, MIME type: image/png)
| This is a file from the Wikimedia Commons. Information from its description page there is shown below. Commons is a freely licensed media file repository. You can help. |
Summary
| DescriptionQuarter of Arthrobacter arilaitensis Re117 genome.png |
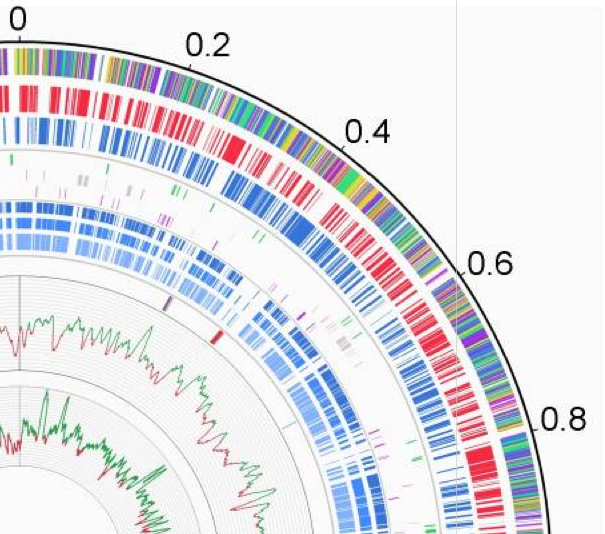
English: The outermost circle (circle 1) represents the scale in Mbp. Circle 2 represents the functional category of the CDSs: green, cell envelope and related processes; blue, intermediary metabolism; yellow, information pathways; orange, other functions; dark purple, proteins of unknown function that are similar to other proteins; light purple, proteins of unknown function without similarity to other proteins. Circles 3 and 4 represent CDSs (excluding transposases) on positive (red) and negative (blue) strands. Circle 5 represents the insertion sequences (in green). Circle 6 represents rRNA (grey) and tRNA (red) genes. Circle 7 represents the pseudogenes (excluding transposase pseudogenes). Circles 8, 9 and 10 represent CDSs (in blue) orthologs (predicted as defined in section “Genome analysis and annotation”) with genes from three environmental Arthrobacter strains: A. chlorophenolicus A6 (circle 8), Arthrobacter sp. FB24 (circle 9) and A. aurescens TC1 (circle 10). Circle 11 represents genes involved in siderophore biosynthesis and export (blue) and in Fe3+/siderophore complexes import (red). Circle 12 and 13 represent the G+C content and GC skew (G−C)/(G+C), respectively, each plotted using a 10-kb window. |
| Date | |
| Source | Modified from Figure 1 in Monnet C, Loux V, Gibrat J-F, Spinnler E, Barbe V, et al. (2010) The Arthrobacter arilaitensis Re117 Genome Sequence Reveals Its Genetic Adaptation to the Surface of Cheese. PLoS ONE 5(11): e15489. doi:10.1371/journal.pone.0015489 |
| Author | C Monnet, V Loux, JF Gibrat, et al |
Licensing
This file is licensed under the Creative Commons Attribution 2.5 Generic license.
- You are free:
- to share – to copy, distribute and transmit the work
- to remix – to adapt the work
- Under the following conditions:
- attribution – You must give appropriate credit, provide a link to the license, and indicate if changes were made. You may do so in any reasonable manner, but not in any way that suggests the licensor endorses you or your use.
Captions
Add a one-line explanation of what this file represents
Items portrayed in this file
depicts
14 June 2011
image/png
File history
Click on a date/time to view the file as it appeared at that time.
| Date/Time | Thumbnail | Dimensions | User | Comment | |
|---|---|---|---|---|---|
| current | 11:57, 14 June 2011 |  | 605 × 534 (432 KB) | Rockpocket |
File usage
More than 100 pages use this file. The following list shows the first 100 pages that use this file only. A full list is available.
- Talk:AMBER
- Talk:Baum–Welch algorithm
- Talk:BioPerl
- Talk:Biocomplexity Institute of Virginia Tech
- Talk:Bioinformatics
- Talk:Biological database
- Talk:Burrows–Wheeler transform
- Talk:C. H. Waddington
- Talk:Cellular automaton
- Talk:Chemical database
- Talk:Clade
- Talk:Cladistics
- Talk:Computational neuroscience
- Talk:Conserved sequence
- Talk:Crossover (genetic algorithm)
- Talk:DNA barcoding
- Talk:DNA microarray
- Talk:Distance matrix
- Talk:Docking (molecular)
- Talk:Dynamic energy budget theory
- Talk:Dynamic programming
- Talk:Ensembl genome database project
- Talk:Eugene Koonin
- Talk:Evolutionary tree
- Talk:FASTA
- Talk:FASTA format
- Talk:Flux balance analysis
- Talk:Folding@home
- Talk:GDB Human Genome Database
- Talk:GenBank
- Talk:Gene prediction
- Talk:Genetic programming
- Talk:Genome
- Talk:Genomics
- Talk:Global Infectious Disease Epidemiology Network
- Talk:Glycomics
- Talk:Haldane's dilemma
- Talk:Hidden Markov model
- Talk:John Maynard Smith
- Talk:Junk DNA
- Talk:Knowledge engineering
- Talk:Lipidomics
- Talk:List of algorithms
- Talk:Long branch attraction
- Talk:Ludwig von Bertalanffy
- Talk:Macromolecular docking
- Talk:Mathematical and theoretical biology
- Talk:Mathematical modelling of infectious diseases
- Talk:Maximum parsimony (phylogenetics)
- Talk:Metabolic network modelling
- Talk:Metabolomics
- Talk:Metagenomics
- Talk:Molecular phylogenetics
- Talk:Most recent common ancestor
- Talk:Mutation (genetic algorithm)
- Talk:Needleman–Wunsch algorithm
- Talk:Neighbor joining
- Talk:Omics
- Talk:Ontology (information science)
- Talk:Petri net
- Talk:Phenome
- Talk:Phylogenetic tree
- Talk:Phylogenetics
- Talk:Phylogeny
- Talk:Point accepted mutation
- Talk:Protein Data Bank
- Talk:Protein structure prediction
- Talk:Protein–protein interaction
- Talk:Proteome
- Talk:Proteomics
- Talk:PubChem
- Talk:PubMed Central
- Talk:PyMOL
- Talk:Regulome
- Talk:Representative sequences
- Talk:Robert Rosen (biologist)
- Talk:Sequence alignment
- Talk:Sequence assembly
- Talk:Sequence clustering
- Talk:Spurious relationship
- Talk:Structural genomics
- Talk:Substitution matrix
- Talk:Synthetic biology
- Talk:Systems biology
- Talk:Systems theory
- Talk:Theoretical ecology
- Talk:Threading (protein sequence)
- Talk:UCPH Bioinformatics Centre
- Talk:Weasel program
- Talk:What Is Life?
- Talk:Wikispecies
- Talk:World Community Grid
- User:Klortho
- User talk:CuriousOliver
- User talk:DarTar
- User talk:Lenov
- User talk:Peak Freak
- User talk:Rockpocket
- User talk:Spitshine
- User talk:Thorwald
View more links to this file.
Global file usage
The following other wikis use this file:
- Usage on fr.wikipedia.org
- Usage on outreach.wikimedia.org