
Enolase
An enzyme involved in glycolysis / From Wikipedia, the free encyclopedia
Dear Wikiwand AI, let's keep it short by simply answering these key questions:
Can you list the top facts and stats about Phosphopyruvate hydratase?
Summarize this article for a 10 year old
Phosphopyruvate hydratase, usually known as enolase, is a metalloenzyme (EC 4.2.1.11) that catalyses the conversion of 2-phosphoglycerate (2-PG) to phosphoenolpyruvate (PEP), the ninth and penultimate step of glycolysis. The chemical reaction is:
- 2-phospho-D-glycerate
phosphoenolpyruvate + H2O
| phosphopyruvate hydratase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Yeast enolase dimer.[1] | |||||||||
| Identifiers | |||||||||
| EC no. | 4.2.1.11 | ||||||||
| CAS no. | 9014-08-8 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Enolase, N-terminal domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
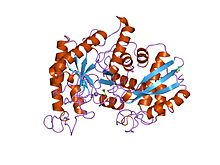 x-ray structure and catalytic mechanism of lobster enolase | |||||||||
| Identifiers | |||||||||
| Symbol | Enolase_N | ||||||||
| Pfam | PF03952 | ||||||||
| Pfam clan | CL0227 | ||||||||
| InterPro | IPR020811 | ||||||||
| PROSITE | PDOC00148 | ||||||||
| SCOP2 | 1els / SCOPe / SUPFAM | ||||||||
| |||||||||
| Enolase | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||
| Identifiers | |||||||||||
| Symbol | Enolase | ||||||||||
| Pfam | PF00113 | ||||||||||
| InterPro | IPR000941 | ||||||||||
| PROSITE | PDOC00148 | ||||||||||
| |||||||||||
Phosphopyruvate hydratase belongs to the family of lyases, specifically the hydro-lyases, which cleave carbon-oxygen bonds. The systematic name of this enzyme is 2-phospho-D-glycerate hydro-lyase (phosphoenolpyruvate-forming).
The reaction is reversible, depending on environmental concentrations of substrates.[3] The optimum pH for the human enzyme is 6.5.[4] Enolase is present in all tissues and organisms capable of glycolysis or fermentation. The enzyme was discovered by Lohmann and Meyerhof in 1934,[5] and has since been isolated from a variety of sources including human muscle and erythrocytes.[4] In humans, deficiency of ENO1 is linked to hereditary haemolytic anemia, while ENO3 deficiency is linked to glycogen storage disease type XIII.
